The process of evolutionary sequence analysis is hindered by both software complexity and the multi-step nature of data integration, management and visualisation. A common workflow will invoke numerous programs and custom scripts, passing information in flat files that are named arbitrarily and can be accidentally overwritten or deleted. This approach works fine for many smaller problems, but does not scale. In practical settings, analysis is performed on data from multiple sources, by multiple collaborators and repeated multiple times with different program options, to integrate new data and update reference information.
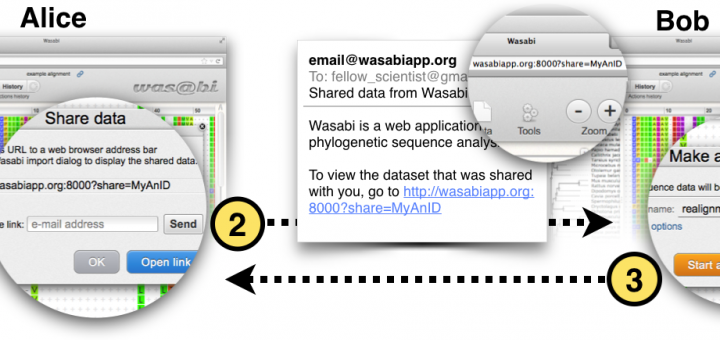
Wasabi is an open source, web–based, integrated environment for evolutionary sequence analysis. Wasabi is built around the phylogeny-aware sequence alignment tools PRANK and PAGAN, which have been shown to produce alignments that better represent evolutionary homology and therefore aid downstream analysis such as inference of selection and phylogenetic tree estimation. Wasabi’s graphical interface simplifies analysis byhiding extraneous options, provides context sensitive menus and integrates with Ensembl to easily allow inclusion of gene sequences, gene family alignments or slices of genomic alignments into projects. Wasabi supports reproducibility by automatically storing the results of intermediate steps and was designed with collaboration and sharing in mind, featuring built–in functions to share data between users and publish analysis results to a wide audience. Wasabi is available at http://wasabiapp.org.